Exercise 1
Exercise 2
Q & A
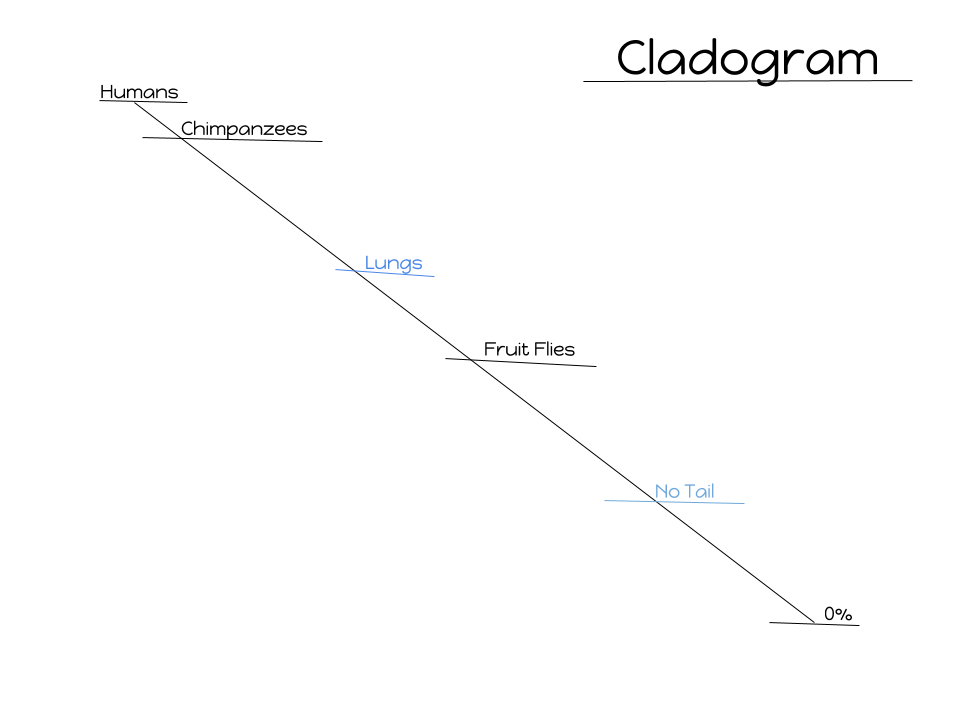
Question 1: According to the cladogram, what organisms have hair?
Answer 1: According to my analyzation of this complex cladogram, the organisms that carry the trait of having hair is the gorilla and tiger.
Question 2: According to the cladogram, what four structures do tigers posses?
Answer 2: The four structures that tigers possess are having jaws, lungs, dry skin, and hair.
Question 3: According to the cladogram, which structure evolved first between lungs and dry skin?
Answer 3: On the cladogram it shows that lungs evolved before dry skin did.
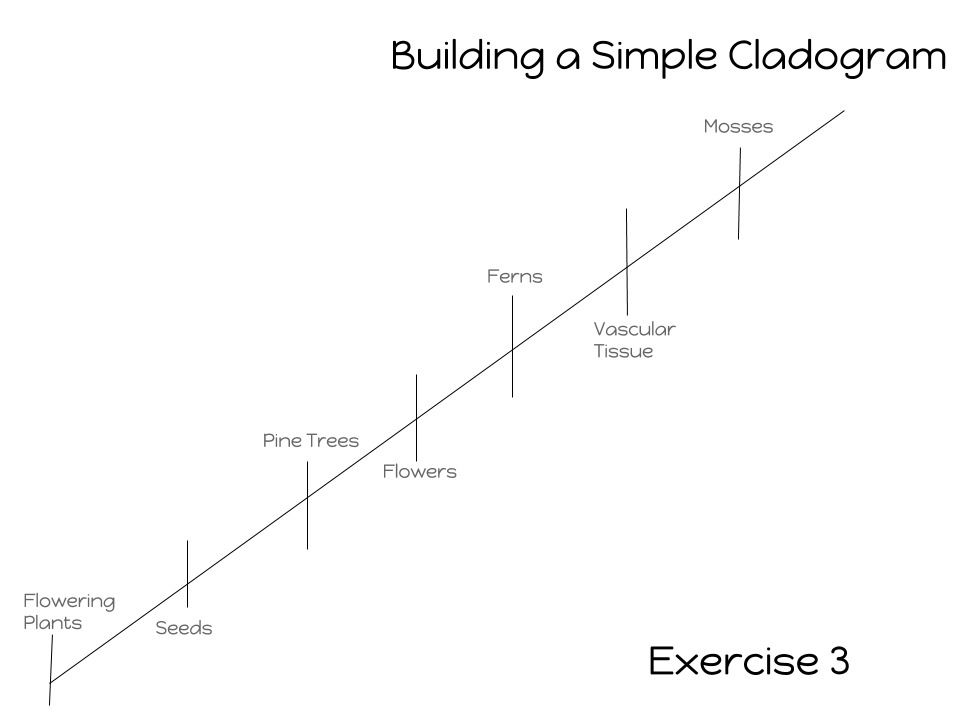
Investigation 2: Building Simple Cladograms
Exercise 3
Exercise 4
A) Why is the percentage of similarity in the protein always higher than the percentage of similarity in the gene for each of the species?
Answer: Proteins are very specific variables and the order of the amino acid is what determines the shape and job of a protein. You can also end up with the sam amino acid even if gene codes are different, and there are 20 amino acids and 64 gene combinations which causes the common protein percentage to be much higher then the common gene percentage.
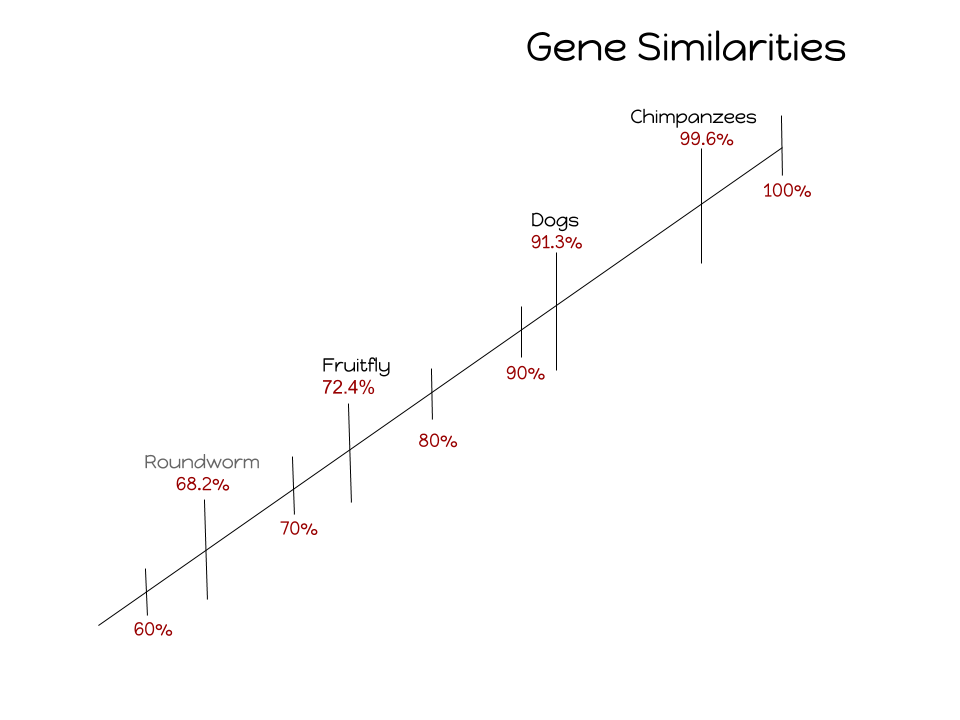
Inserted below are the percentage of gene similarities between al five species
Answer: Proteins are very specific variables and the order of the amino acid is what determines the shape and job of a protein. You can also end up with the sam amino acid even if gene codes are different, and there are 20 amino acids and 64 gene combinations which causes the common protein percentage to be much higher then the common gene percentage.
Inserted below are the percentage of gene similarities between al five species
Investigation 3: Uncovering Fossil Specimen Using BLAST
Uncovering Fossil Specimen Using BLAST
A. Morphological Observation of the Fossil Specimen and Forming the Hypothesis
Observations
~ Spine leading up to the end of its tail
~ Two thin short legs
~ A dark round eye
~ A hald circled face
~ Ribs
~ Very short neck
~ Spine leading up to the end of its tail
~ Two thin short legs
~ A dark round eye
~ A hald circled face
~ Ribs
~ Very short neck
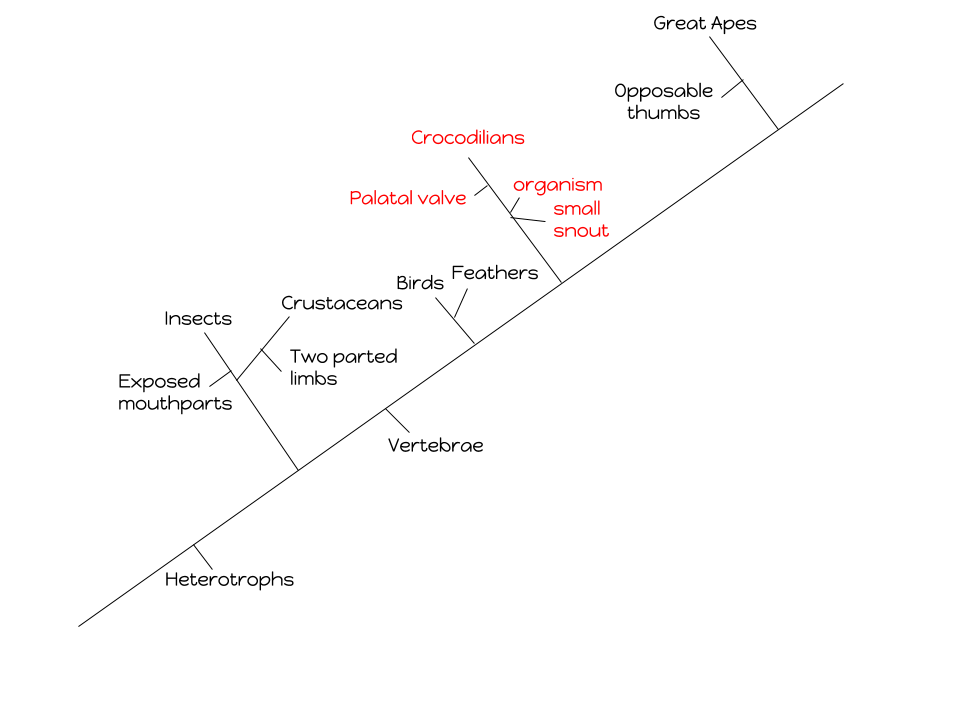
Hypothesis
If the unidentified organism has a tail, then it will most likely fall under the same category as the Crocodilians.
If the unidentified organism has a tail, then it will most likely fall under the same category as the Crocodilians.
Using Blast to Analyze Genes and Determine the Most Likely Placement of the Fossil Species
Step 1: Instruction for Downloading Gene Files
Before starting this investigation, students need to download four gene files (Gene 1-Gene 4) to their fl ash drive or computer. These files are located at the following web address: http://blogging4biology.edublogs.org/2010/08/28/college-board-lab-fi les/
Note that these files will not open on your computer. They only work when opened on the BLAST website.
Step 2: Instructions for BLAST Queries
Upload the gene sequences into BLAST by following the instructions below:
1. Go to the BLAST homepage: http://blast.ncbi.nlm.nih.gov/Blast.cgi
2. Click on “Saved Strategies” on the top of the page.
3. Under “Upload Search Strategy,” click on “Browse” and locate one of the gene files you saved onto your computer.
4. Click “View.”
5. A screen will appear with the parameters for your query already configured.
Note Do not alter any of the parameters. Scroll down the page and click on the “BLAST” button at the bottom.
6. After collecting and analyzing all of the data for that particular gene (see instructions below), repeat this procedure for the other three gene sequences.
Step 3: Instructions for Analyzing BLAST Queries
~ Scroll down to the section titled, “Sequences producing significant alignments”. The list of organisms that appear below this section are those with sequences identical to or most similar to the gene of interest. The
most similar sequences are listed first and as you move down the list, the sequences become less similar to your gene of interest.
~ Try clicking on a particular species listed to get a full report that includes the species’ classification scheme, the research journal the gene was first reported in, and the sequence of bases that appear to align with your gene of interest.
~ Click “Distance tree of results,” a cladogram of the species with similar sequences to your gene of interest placed on the cladogram will be shown according to how closely their matched gene aligns with your gene of interest.
Step 4: Analysis of Results
~ Species share similar genes because of common ancestry. The more similar genes two species have in common, the more recent their common ancestor. Thus, the two species will be located closer on a cladogram.
~ As you collect information from BLAST for each of the gene files explain whether the data supports your original hypothesis and your original placement of the fossil species on the cladogram.
Gene File 1
What species has the most similar gene sequence as your gene of interest?
~ Gallus Collagen is the most similar gene sequence.
Where is that species located on the cladogram?
~ The species is located directly under gene 1.
How similar is that gene sequence?
~ There is 100% gene sequence similarity.
What species has the least similar gene sequence as your gene interest?
~ Anolis Corolinensis has the least similar gene sequence.
Gene File 2
What species has the most similar gene sequence as your gene of interest?
~ Flies have the most similar gene sequence as my gene.
Where is that species located on the cladogram?
~ The species is 6 species above gene 2
How similar is that gene sequence?
~ There is 100% gene sequence similarity.
What species has the least similar gene sequence as your gene interest?
~ The Pediculus Humanus is the least similar to my gene.
Gene File 3
What species has the most similar gene sequence as your gene of interest?
~ The Taeniopygia Guttata has the most similar gene sequence.
Where is that species located on the cladogram?
~The species is right about gene 3.
How similar is that gene sequence?
~ There is 95% gene sequence similarity.
What species has the most similar gene sequence as your gene of interest?
~ The Sus Scrofa is the least similar to my gene
Gene File 4
What species has the most similar gene sequence as your gene of interest?
~ The alligator has the most similar gene sequence as my gene of interest.
Where is that species located on the cladogram?
~ The alligator is located under gene 4.
How similar is that gene sequence?
~ There is exactly a 100% gene sequence similarity.
What species has the most similar gene sequence as your gene of interest?
~ Barbourbula Busuangensis has the least similar gene sequence.
Drawing A Cladogram
Conclusion
According to my analysis, my previously stated hypothesis is correct. The organism that shares the least amount of gene sequence similarities is the insect, so therefore I decided to place my unknown organism on the same branch that the Crocodilians are located on. Gene 4 is what shares a 100% rate gene similarity sequence with the gene sequence belonging to the alligator.
Investigation 4: Blast Your Own Genes of Interest!
Investigation 4: Blast Your Own Genes of Interest!
1. On the Entrez Gene website search the term “human actin”.
2. Click on the fi rst link that appears and scroll down to “NCBI Reference Sequences”.
3. Click on the fi rst fi le name “NM 001100.3” under “mRNA and Proteins”.
4. Click on “FASTA” just below the gene title.
5. The nucleotide sequence displayed is that of the actin gene in humans.
6. Copy the gene sequence and go to the BLAST homepage (http://blast.ncbi.nlm.nih.gov/Blast.cgi).
7. Click on “nucleotide blast” under the Basic BLAST menu.
8. Paste the sequence into the box where it says “Enter Query Sequence.”
9. Give the query a title in the box provided if you plan on saving it for later.
10. Under “Choose Search Set” select the type or genome you want to search (human genome, mouse genome, or all genomes available).
11. Under “Program Selection” choose whether or not you want highly similar sequences or somewhat similar sequences. Choosing somewhat similar sequences will provide you with more results.
12. Click BLAST.In humans, what is the importance of the gene you chose? Would you expect to find that gene in all organisms? Why or why not?